Backend Module Design
GPJax is built upon Equinox and Paramax. Equinox provides a lightweight module system for JAX, while Paramax adds support for constrained parameters via unwrappable types. This notebook provides a high-level overview of the backend module design in GPJax. For an introduction to Equinox, please refer to the official documentation.
import typing as tp
import equinox as eqx
from examples.utils import use_mpl_style
from gpjax.mean_functions import (
AbstractMeanFunction,
Constant,
)
from gpjax.parameters import (
PositiveReal,
Real,
)
from gpjax.typing import (
Array,
ScalarFloat,
)
# Enable Float64 for more stable matrix inversions.
from jax import config
import jax.numpy as jnp
import jax.tree_util as jtu
from jaxtyping import (
Float,
Num,
install_import_hook,
)
import matplotlib as mpl
import matplotlib.pyplot as plt
import paramax
from paramax import AbstractUnwrappable
config.update("jax_enable_x64", True)
with install_import_hook("gpjax", "beartype.beartype"):
import gpjax as gpx
# set the default style for plotting
use_mpl_style()
cols = mpl.rcParams["axes.prop_cycle"].by_key()["color"]
Parameters
GPJax uses Paramax to handle constrained parameters. As discussed in our Sharp Bits - Bijectors Doc, GPJax uses bijectors to transform constrained parameters to unconstrained parameters during optimisation. You may register the support of a parameter using our parameter types. To see this, consider the constant mean function which contains a single constant parameter whose value ordinarily exists on the real line. We can register this parameter as follows:
Constant(constant=Real(value=weak_f64[]))
However, suppose you wish your mean function's constant parameter to be strictly
positive. This is easy to achieve by using the correct parameter type which, in this
case, will be the PositiveReal. All parameter types are subclasses of Paramax's
AbstractUnwrappable, which means they will be automatically transformed by GPJax
during optimisation.
True
Injecting this newly constrained parameter into our mean function is then identical to before.
Constant(constant=PositiveReal(_unconstrained=weak_f64[]))
Parameter Transforms
With a parameter instantiated, you likely wish to transform the parameter's value from
its constrained support onto the entire real line. In GPJax, parameters store their
values internally in unconstrained space. When you need the constrained value, you
simply call unwrap() on the parameter, or use paramax.unwrap() on an entire model
to resolve all parameters at once.
print("Constrained value:", constant_param.unwrap())
print("Unconstrained (internal) value:", constant_param._unconstrained)
Constrained value: 1.0
Unconstrained (internal) value: 0.5413248546129181
We see here that the Softplus bijector is applied by the PositiveReal parameter type.
Internally, the value 1.0 is stored as its inverse-softplus (~0.54), and calling
unwrap() applies softplus to recover the original constrained value.
For a value closer to 0, the transformation is more pronounced.
close_to_zero_param = PositiveReal(value=1e-6)
print("Constrained value:", close_to_zero_param.unwrap())
print("Unconstrained (internal) value:", close_to_zero_param._unconstrained)
Constrained value: 9.999999999999985e-07
Unconstrained (internal) value: -13.815510057964234
Transforming Multiple Parameters
In the above, we transformed a single parameter. However, in practice your parameters may be nested within several functions e.g., a kernel function within a GP model. Fortunately, transforming several parameters is a simple operation that we here demonstrate for a conjugate GP posterior (see our Regression Notebook for detailed explanation of this model.).
kernel = gpx.kernels.Matern32()
meanf = gpx.mean_functions.Constant()
prior = gpx.gps.Prior(mean_function=meanf, kernel=kernel)
likelihood = gpx.likelihoods.Gaussian(100)
posterior = likelihood * prior
print(posterior)
ConjugatePosterior(
prior=Prior(
kernel=Matern32(
active_dims=slice(None, None, None),
compute_engine=<gpjax.kernels.computations.dense.DenseKernelComputation object at 0x7f8de2707d10>,
lengthscale=PositiveReal(_unconstrained=weak_f64[]),
variance=NonNegativeReal(_unconstrained=weak_f64[])
),
mean_function=Constant(constant=weak_f64[])
),
likelihood=Gaussian(
num_datapoints=100,
integrator=<gpjax.integrators.AnalyticalGaussianIntegrator object at 0x7f8de82a6310>,
obs_stddev=NonNegativeReal(_unconstrained=weak_f64[])
)
)
Now contained within the posterior there are four parameters: the kernel's lengthscale
and variance, the noise variance of the likelihood, and the constant of the mean
function. With Equinox, we can partition the model into its array leaves and static
structure using eqx.partition. This gives us direct access to the parameters as a
PyTree.
ConjugatePosterior(
prior=Prior(
kernel=Matern32(
active_dims=slice(None, None, None),
compute_engine=<gpjax.kernels.computations.dense.DenseKernelComputation object at 0x7f8de2707d10>,
lengthscale=PositiveReal(_unconstrained=weak_f64[]),
variance=NonNegativeReal(_unconstrained=weak_f64[])
),
mean_function=Constant(constant=weak_f64[])
),
likelihood=Gaussian(
num_datapoints=100,
integrator=<gpjax.integrators.AnalyticalGaussianIntegrator object at 0x7f8de82a6310>,
obs_stddev=NonNegativeReal(_unconstrained=weak_f64[])
)
)
The params object behaves just like a PyTree and, consequently, we may use JAX's
tree_map function to alter the values. The updated params can then be recombined
with the static structure using eqx.combine. In the below, we simply increment each
parameter's value by 1.
ConjugatePosterior(
prior=Prior(
kernel=Matern32(
active_dims=slice(None, None, None),
compute_engine=<gpjax.kernels.computations.dense.DenseKernelComputation object at 0x7f8de2707d10>,
lengthscale=PositiveReal(_unconstrained=weak_f64[]),
variance=NonNegativeReal(_unconstrained=weak_f64[])
),
mean_function=Constant(constant=weak_f64[])
),
likelihood=Gaussian(
num_datapoints=100,
integrator=<gpjax.integrators.AnalyticalGaussianIntegrator object at 0x7f8de82a6310>,
obs_stddev=NonNegativeReal(_unconstrained=weak_f64[])
)
)
Let us now use Equinox's combine function to reconstruct the posterior distribution
using the updated parameters.
ConjugatePosterior(
prior=Prior(
kernel=Matern32(
active_dims=slice(None, None, None),
compute_engine=<gpjax.kernels.computations.dense.DenseKernelComputation object at 0x7f8de2707d10>,
lengthscale=PositiveReal(_unconstrained=weak_f64[]),
variance=NonNegativeReal(_unconstrained=weak_f64[])
),
mean_function=Constant(constant=weak_f64[])
),
likelihood=Gaussian(
num_datapoints=100,
integrator=<gpjax.integrators.AnalyticalGaussianIntegrator object at 0x7f8de82a6310>,
obs_stddev=NonNegativeReal(_unconstrained=weak_f64[])
)
)
To resolve all constrained parameter values at once (applying each parameter's
bijection), we can use paramax.unwrap on the entire model.
ConjugatePosterior(
prior=Prior(
kernel=Matern32(
active_dims=slice(None, None, None),
compute_engine=<gpjax.kernels.computations.dense.DenseKernelComputation object at 0x7f8de2707d10>,
lengthscale=weak_f64[],
variance=weak_f64[]
),
mean_function=Constant(constant=weak_f64[])
),
likelihood=Gaussian(
num_datapoints=100,
integrator=<gpjax.integrators.AnalyticalGaussianIntegrator object at 0x7f8de82a6310>,
obs_stddev=weak_f64[]
)
)
Fine-Scale Control
One of the advantages of Equinox's partition mechanism is that we can gain fine-scale
control over which parameters we extract. For example, suppose we only wish to extract
those parameters whose support is the positive real line. This is easily achieved by
providing a custom filter function to eqx.partition.
positive_reals, other_params = eqx.partition(
posterior, lambda leaf: isinstance(leaf, PositiveReal)
)
print(positive_reals)
ConjugatePosterior(
prior=Prior(
kernel=Matern32(
active_dims=slice(None, None, None),
compute_engine=<gpjax.kernels.computations.dense.DenseKernelComputation object at 0x7f8de2707d10>,
lengthscale=PositiveReal(_unconstrained=None),
variance=NonNegativeReal(_unconstrained=None)
),
mean_function=Constant(constant=None)
),
likelihood=Gaussian(
num_datapoints=100,
integrator=<gpjax.integrators.AnalyticalGaussianIntegrator object at 0x7f8de82a6310>,
obs_stddev=NonNegativeReal(_unconstrained=None)
)
)
Now we see that we have two objects: one containing the positive real parameters and the other containing the remaining structure. This functionality is exceptionally useful as it allows us to efficiently operate on a subset of the parameters whilst leaving the others untouched. Looking forward, we hope to use this functionality in our Variational Inference Approximations to perform more efficient updates of the variational parameters and then the model's hyperparameters.
Equinox Modules
To conclude this notebook, we will now demonstrate the ease of use and flexibility offered by Equinox modules. To do this, we will implement a linear mean function using the existing abstractions in GPJax.
For inputs \(x_n \in \mathbb{R}^d\), the linear mean function \(m(x): \mathbb{R}^d \to \mathbb{R}\) is defined as: $$ m(x) = \alpha + \sum_{i=1}^d \beta_i x_i $$ where \(\alpha \in \mathbb{R}\) and \(\beta_i \in \mathbb{R}\) are the parameters of the mean function. Let's now implement that using Equinox.
class LinearMeanFunction(AbstractMeanFunction):
intercept: Real | Float[Array, " O"]
slope: Real | Float[Array, " D O"]
def __init__(
self,
intercept: ScalarFloat | Float[Array, " O"] | Real = 0.0,
slope: ScalarFloat | Float[Array, " D O"] | Real = 0.0,
):
if isinstance(intercept, Real):
self.intercept = intercept
else:
self.intercept = Real(jnp.array(intercept))
if isinstance(slope, Real):
self.slope = slope
else:
self.slope = Real(jnp.array(slope))
def __call__(self, x: Num[Array, "N D"]) -> Float[Array, "N O"]:
# Use a helper that works whether the parameter is still wrapped
# (an AbstractUnwrappable) or has already been unwrapped to a plain
# array by paramax.unwrap().
def _val(p):
return p.unwrap() if isinstance(p, AbstractUnwrappable) else p
return _val(self.intercept) + jnp.dot(x, _val(self.slope))
As we can see, the implementation is straightforward and concise. The
AbstractMeanFunction is a subclass of eqx.Module and may, therefore, be
used in any partition or combine call. Further, we have registered the intercept
and slope parameters as Real parameter types. This registers their value in the
PyTree and means that they will be part of any operation applied to the model e.g.,
unwrapping and differentiation.
To check our implementation worked, let's now plot the value of our mean function for a linearly spaced set of inputs.
N = 100
X = jnp.linspace(-5.0, 5.0, N)[:, None]
meanf = LinearMeanFunction(intercept=1.0, slope=2.0)
plt.plot(X, meanf(X))
[<matplotlib.lines.Line2D at 0x7f8d97b2e650>]
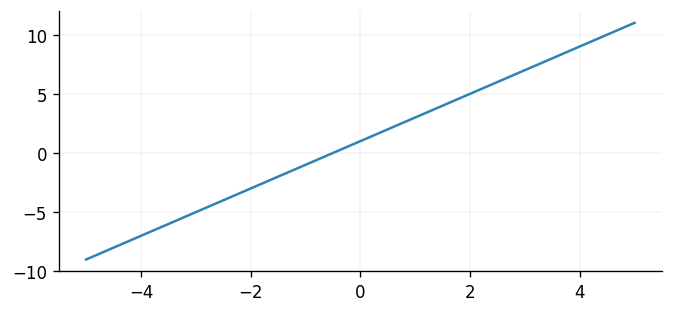
Looks good! To conclude this section, let's now parameterise a GP with our new mean function and see how gradients may be computed.
y = jnp.sin(X)
D = gpx.Dataset(X, y)
prior = gpx.gps.Prior(mean_function=meanf, kernel=gpx.kernels.Matern32())
likelihood = gpx.likelihoods.Gaussian(D.n)
posterior = likelihood * prior
We'll compute derivatives of the conjugate marginal log-likelihood. With Equinox and
Paramax, this is straightforward: paramax.unwrap resolves all constrained parameters
inside the loss function, and eqx.filter_value_and_grad computes gradients with
respect to the array leaves of the model.
def loss_fn(model, data: gpx.Dataset) -> ScalarFloat:
model = paramax.unwrap(model)
return -gpx.objectives.conjugate_mll(model, data)
_, param_grads = eqx.filter_value_and_grad(loss_fn)(posterior, D)
In practice, you would wish to perform multiple iterations of gradient descent to
learn the optimal parameter values. However, for the purposes of illustration, we use
eqx.apply_updates in the below to update the model using its previously computed
gradients. As you can see, Equinox makes it easy to apply updates directly to the
model without manual split/merge operations.
LEARNING_RATE = 0.01
scaled_grads = jtu.tree_map(lambda g: LEARNING_RATE * g, param_grads)
optimised_posterior = eqx.apply_updates(posterior, scaled_grads)
Now we will plot the updated mean function alongside its initial form. Since the model
is updated in-place via eqx.apply_updates, we can simply invoke it as normal.
fig, ax = plt.subplots()
ax.plot(X, optimised_posterior.prior.mean_function(X), label="Updated mean function")
ax.plot(X, meanf(X), label="Initial mean function")
ax.legend()
ax.set(xlabel="x", ylabel="m(x)")
[Text(0.5, 0, 'x'), Text(0, 0.5, 'm(x)')]
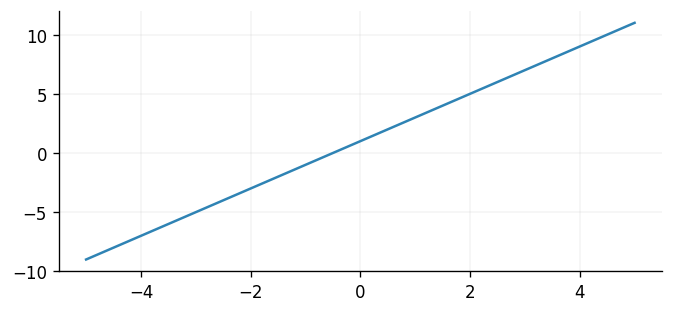
Conclusions
In this notebook we have explored how GPJax's Equinox-based backend may be easily manipulated and extended. For a more applied look at this, see how we construct a kernel on polar coordinates in our Kernel Guide notebook.
System configuration
Author: Thomas Pinder
Last updated: Mon, 25 May 2026
Python implementation: CPython
Python version : 3.11.15
IPython version : 9.9.0
equinox : 0.13.5
gpjax : 0.14.0
jax : 0.9.0
jaxtyping : 0.3.6
matplotlib: 3.10.8
paramax : 0.0.5
Watermark: 2.6.0