Joint Inference with Numpyro
In this notebook, we demonstrate how to use Numpyro to perform fully Bayesian inference over the hyperparameters of a Gaussian process model. We will look at a scenario where we have a structured mean function in the form of a linear model, and a GP capturing the residuals. We will infer the parameters of both the linear model and the GP jointly.
from examples.utils import use_mpl_style
import gpjax as gpx
from jax import config
import jax.numpy as jnp
import jax.random as jr
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpyro
import numpyro.distributions as dist
from numpyro.infer import (
MCMC,
NUTS,
Predictive,
)
config.update("jax_enable_x64", True)
use_mpl_style()
cols = mpl.rcParams["axes.prop_cycle"].by_key()["color"]
key = jr.key(123)
keys = jr.split(key, 4)
Data Generation
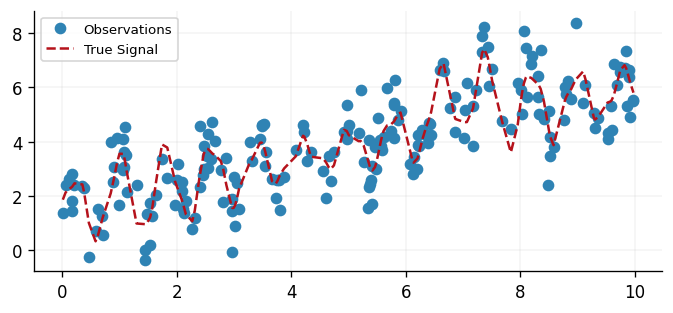
We generate a synthetic dataset that consists of a linear trend together with a locally periodic residual signal whose amplitude varies over time, an additional high-frequency component, and a local bump. This data generating process is purposefully designed to illustrate the benefit of incorporating a Gaussian process into a larger Bayesian model; however, such structures are common.
N = 200
x = jnp.sort(jr.uniform(keys[0], shape=(N, 1), minval=0.0, maxval=10.0), axis=0)
# True parameters for the linear trend
true_slope = 0.45
true_intercept = 1.5
# Structured residual signal captured by the GP
slow_period = 6.0
fast_period = 0.8
amplitude_envelope = 1.0 + 0.5 * jnp.sin(2 * jnp.pi * x / slow_period)
modulated_periodic = amplitude_envelope * jnp.sin(2 * jnp.pi * x / fast_period)
high_frequency_component = 0.3 * jnp.cos(2 * jnp.pi * x / 0.35)
localised_bump = 1.2 * jnp.exp(-0.5 * ((x - 7.0) / 0.45) ** 2)
linear_trend = true_slope * x + true_intercept
residual_signal = modulated_periodic + high_frequency_component + localised_bump
signal = linear_trend + residual_signal
# Observations with homoscedastic noise
observation_noise = 0.7
y = signal + observation_noise * jr.normal(keys[1], shape=x.shape)
D = gpx.Dataset(X=x, y=y)
fig, ax = plt.subplots()
ax.plot(x, y, "o", label="Observations", color=cols[0])
ax.plot(x, signal, "--", label="True Signal", color=cols[1])
ax.legend()
<matplotlib.legend.Legend at 0x7f35a506d1d0>
Model Definition
We define a GP model with a zero mean function, as we will handle the linear trend explicitly in the Numpyro model. Naturally, one could parameterise the GP with a linear mean function; however, this design is purely pedagogical. For the kernel, we specify a product of a periodic kernel and an RBF kernel. This choice reflects our prior knowledge that the signal is locally periodic. For a more in-depth look at how complex kernels can be designed, see our Introduction to Kernels notebook.
We see in the below that priors are specified on the parameters' constrained space. For
example, the lengthscale parameter must be strictly positive and, therefore, a unit-Gaussian
would be a poor choice of prior. Instead, we opt for the log-Gaussian as the prior distribution
as its support matches that of our lengthscale parameter. Priors are standard
NumPyro distributions sampled directly
inside the model function with numpyro.sample.
# Priors are defined as NumPyro distributions and sampled directly inside the model
# function below. GPJax parameter constructors accept raw JAX arrays from
# numpyro.sample, so no special registration step is needed.
We'll construct the Gaussian process inside the NumPyro model function, passing sampled hyperparameters directly to the GPJax constructors. For a deeper look at how GP construction works, see our Regression notebook.
Joint Inference Loop
We define a NumPyro model that samples all parameters directly using
numpyro.sample, builds the GPJax posterior from those samples, and
scores it with the conjugate marginal log-likelihood via numpyro.factor.
No special registration step is needed -- GPJax constructors accept raw
JAX arrays returned by numpyro.sample.
def model(X, Y, X_new=None):
slope = numpyro.sample("slope", dist.Normal(0.0, 2.0))
intercept = numpyro.sample("intercept", dist.Normal(0.0, 2.0))
linear_component = slope * X + intercept
residuals = Y - linear_component
lengthscale = numpyro.sample("lengthscale", dist.LogNormal(0.0, 1.0))
variance = numpyro.sample("variance", dist.LogNormal(0.0, 1.0))
period = numpyro.sample("period", dist.LogNormal(0.0, 0.5))
obs_noise = numpyro.sample("obs_noise", dist.LogNormal(0.0, 1.0))
stationary_component = gpx.kernels.RBF(
lengthscale=lengthscale, variance=variance
)
periodic_component = gpx.kernels.Periodic(
lengthscale=lengthscale, period=period
)
kernel = stationary_component * periodic_component
meanf = gpx.mean_functions.Constant()
prior = gpx.gps.Prior(mean_function=meanf, kernel=kernel)
likelihood = gpx.likelihoods.Gaussian(num_datapoints=N, obs_stddev=obs_noise)
posterior = prior * likelihood
D_resid = gpx.Dataset(X=X, y=residuals)
mll = gpx.objectives.conjugate_mll(posterior, D_resid)
numpyro.factor("gp_log_lik", mll)
if X_new is not None:
latent_dist = posterior.predict(X_new, train_data=D_resid)
f_new = numpyro.sample("f_new", latent_dist)
f_new = f_new.reshape((-1, 1))
y_noise = numpyro.sample(
"y_noise",
dist.Normal(0.0, obs_noise).expand(f_new.shape).to_event(f_new.ndim),
)
total_prediction = slope * X_new + intercept + f_new + y_noise
numpyro.deterministic("y_pred", total_prediction)
return total_prediction
Running MCMC
Using Numpyro's NUTS sampler, we can now draw samples from the posterior. To ensure
our documentation can be quickly built, we limit the number of samples and the length
of the burn-in phase below. However, in practice, one should draw more samples from
multiple chains using the num_chains argument in the MCMC constructor.
nuts_kernel = NUTS(model)
# In practice, one should run more samples from multiple chains.
mcmc = MCMC(
nuts_kernel,
num_warmup=500,
num_samples=500,
)
mcmc.run(keys[2], x, y)
mcmc.print_summary()
mean std median 5.0% 95.0% n_eff r_hat
intercept 0.59 1.45 0.60 -1.97 2.67 440.39 1.00
lengthscale 2.43 0.75 2.31 1.31 3.53 277.04 1.00
obs_noise 0.79 0.04 0.79 0.72 0.86 380.26 1.00
period 0.80 0.02 0.80 0.77 0.83 210.58 1.01
slope 0.51 0.29 0.50 0.03 0.97 303.14 1.00
variance 7.46 5.01 5.86 1.58 15.07 309.37 1.00
Number of divergences: 0
Analysis and Plotting
Having obtained samples from the posterior, we now evaluate the predictive posterior
at the test sites. In our
Poisson Regression, this
process is done manually. However, by virtue of using Numpyro here, we may instead
use Numpyro's Predictive object to handle this process for us. Once samples are
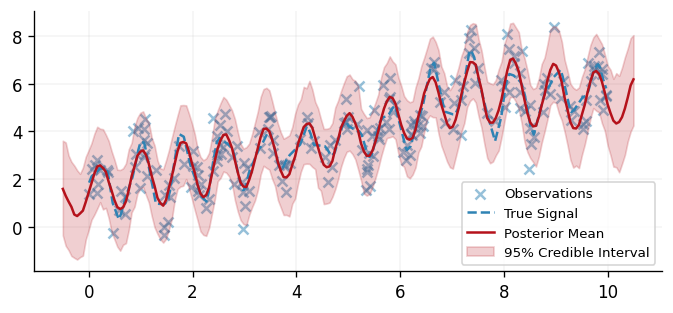
drawn from the predictive posterior distribution, we may evaluate the mean and 95%
credible interval and compare our model's predictions to the underlying data.
samples = mcmc.get_samples()
predictive = Predictive(
model,
posterior_samples=samples,
return_sites=["y_pred"],
)
x_test = jnp.linspace(-0.5, 10.5, 200).reshape(-1, 1)
predictions = predictive(keys[3], x, y, X_new=x_test)
y_pred = predictions["y_pred"]
mean_prediction = jnp.mean(y_pred, axis=0)
lower, upper = jnp.percentile(y_pred, jnp.array([2.5, 97.5]), axis=0)
fig, ax = plt.subplots()
ax.scatter(x, y, alpha=0.5, label="Observations", color=cols[0])
ax.plot(x, signal, "--", label="True Signal", color=cols[0])
ax.plot(x_test, mean_prediction, "-", label="Posterior Mean", color=cols[1])
ax.fill_between(
x_test.flatten(),
lower.flatten(),
upper.flatten(),
color=cols[1],
alpha=0.2,
label="95% Credible Interval",
)
ax.legend()
<matplotlib.legend.Legend at 0x7f359ce9b1d0>
Conclusions
This concludes our introduction to the integration of GPJax with Numpyro. The presentation given here is designed to best illustrate how the two libraries integrate. For a closer look at the more complex models that one may build by integrating Numpyro and GPJax, see our Spatial Semi-Linear Model notebook.
System configuration
Author: Thomas Pinder
Last updated: Mon, 18 May 2026
Python implementation: CPython
Python version : 3.11.15
IPython version : 9.9.0
gpjax : 0.14.0
jax : 0.9.0
matplotlib: 3.10.8
numpyro : 0.19.0
Watermark: 2.6.0
We currently have some availability for consulting on how Gaussian processes, Bayesian modelling, and GPJax can be integrated into your team's work. If this sounds relevant to your work, book an introductory call. These calls are for consulting inquiries only. For technical usage questions and free community support, please use GitHub Discussions and the documentation below.